A better understanding of the relationship between the genes could aid in developing prognostic markers and better therapeutic strategies in cancer treatment. One example that may be of interest to proteomics researchers is Cytoscape.17 Cytoscape is a bioinformatics software platform for visualizing molecular. Most of the genes mainly participated in gene and protein expression, cell signaling and metabolism. Biological Networks Gene Ontology (BiNGO) was used to determine two ontologies (molecular function and biological process) involved in the protein network. We found eight specific non-overlapping clusters of various sizes, which emerged from the huge network of protein-interactors using MCODE version 1.32 clustering algorithm. The 1762 protein interactions were assembled and visualized in y organic form. A total 500 genes from STRING database loaded into Cytoscape version 2.8.3. as a plugin for Cytoscape, which is a an open source bioinformatics software. UCINET: Software for social network analysis that includes a variety of tools and algorithms for analyzing social networks. Current version : BiNGO 3.0.3 (compatible with Cytoscape 3.0 and above). High-throughput experimental data, literature data and hypothetical studies have been used to determine the roles of candidate genes in TP53 pathway. Introduction to Social Network Methods: A comprehensive online textbook that introduces the basics of social network analysis. Its most common use case is as a visualization software component, so it can be used to. TP53 protein interaction database was obtained from STRING version 9.1 program. Cytoscape.js is an open-source JavaScript-based graph library. 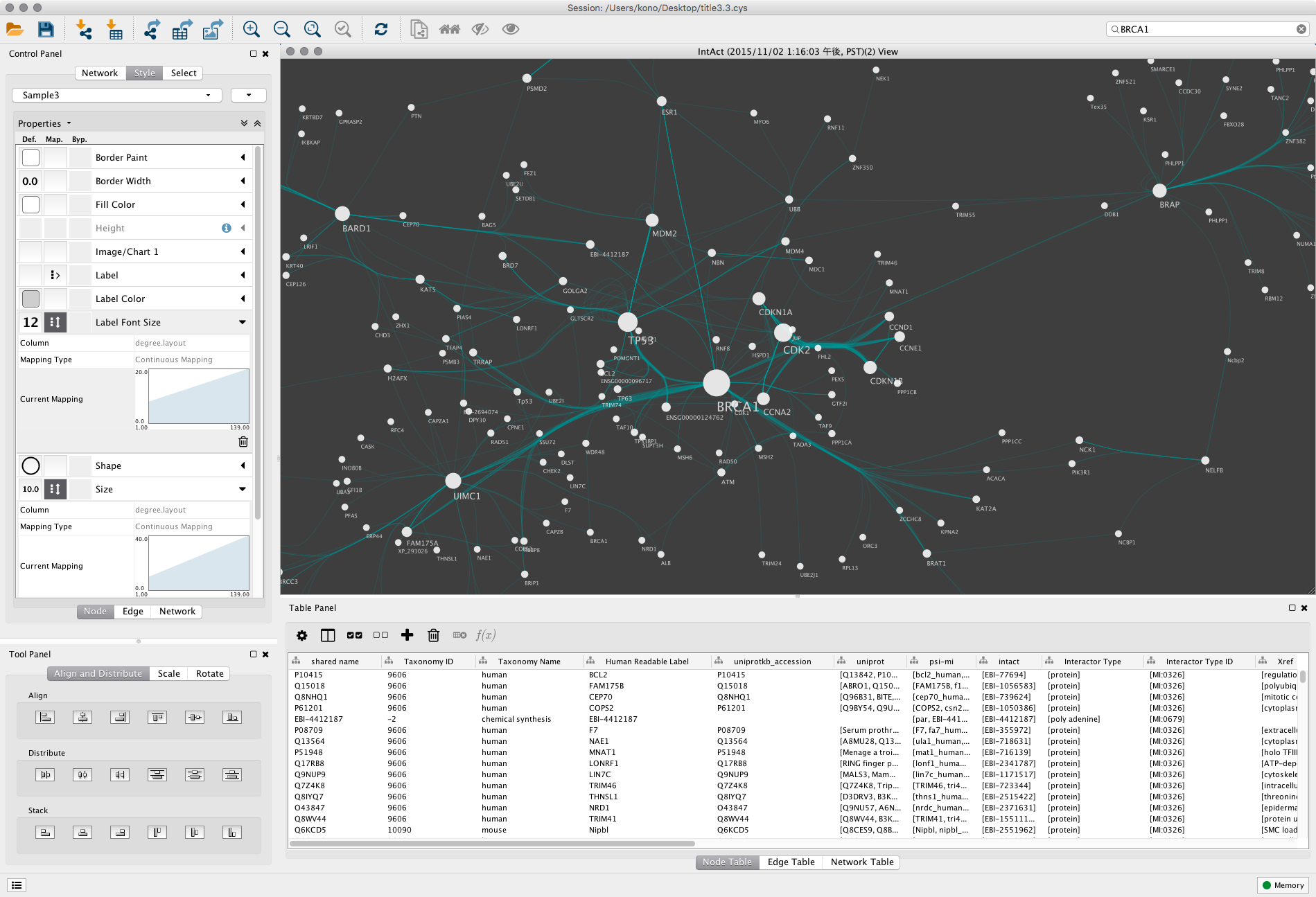
p53 gene alteration could cause uncontrolled cell proliferation.In the present study, we used TP53 gene as the seed in the construction of a protein-protein functional association network to identify genes that might involve in tumorgenesis process with TP53. It induces cell cycle arrest or apoptosis in response to cellular stress and damage. Cytoscape is a Java application verified to run on the Linux, Windows, and Mac OS X platforms. Its core is to provide a basic functional layout and query network. In addition, Cytoscape has an open architecture so that others can extend. When users first install Cytoscape, it has a small well-defined set of functionality for working with networks. TP53 acts as a tumor suppressor in cancer. Cytoscape is a software that focuses on open-source network visualization and analysis. Cytoscape is an open source software tool for network visualization and analysis, primarily for biological data ( Cline et al., 2007 Shannon et al., 2003 ). 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
